Applying Complex Models on Genomic Data
by Patrick Durand, Dominique Lavenier, Michel Leborgne, Anne Siegel, Philippe Veber and Jacques Nicolas
The Symbiose team at IRISA-INRIA is involved in large-scale genome studies using complex models. These encompass genome modelling, development of dedicated hardware and gene networks modelling. Symbiose is also in charge of the bioinformatics platform of the OUEST-genopole research structure.
Symbiose is an INRIA project in bioinformatics, and comprises 25 people. Our research includes large-scale genome studies and complex pattern filtering methods. Principal areas of focus are genome modelling with formal languages, development of dedicated machines and gene networks modelling. Applications include the discovery of target proteins, the study of mutations of toxic bacteria such as Staphylococcus aureus, the prediction of disulfide bonds involved in processes such as protein aggregation, high-speed searching of huge databases and signalling of TGF-beta in liver cancer.
More Expressive Models on Sequences
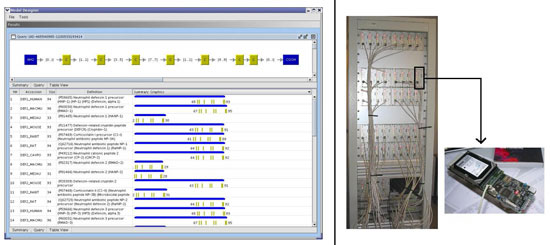
Locating sequences of medical interest in genomic databases can be efficiently performed using a sequence model (also called a pattern). Such a model describes a region of biological interest characteristic of a family of sequences using various classes of languages such as regular expressions, definite clause grammars and string variable grammars (SVG). These provide a high degree of expressiveness, allowing the modelling of high-order sequence organization (such as copies, palindromes or even structural properties like stem-loop or pseudo-knots). In this context, we have developed STAN, a piece of software that provides the researcher with a SVG-based search system. To perform efficient scans, STAN uses a suffix-tree data structure. Since the usage of SVG syntax is quite difficult, we have also designed ModelDesigner, a graphical software platform that allows the user to create a model graphically, without the need for a programming language.
An Application in Anti-Microbial Discovery
In collaboration with INSERM U435 laboratory, the Symbiose team scanned the human genome to search for new anti-microbial peptides known as defensins. These promising compounds are components of innate immunity that are produced naturally by vertebrates. We first defined a model of the genes in question, by extracting the characteristic signature of genes in this family from known defensin sequences. The human genome was then scanned with the translation of the protein model. The STAN and ModelDesigner software previously described found forty new human defensins, of which thirteen putative defensins have now been synthesized and tested with success. This research is leading to the development of a spin-off biotechnology company.
More Computing Power for Genomic Applications
Genomic databases nearly double every year. Applying complex models on this huge data set is a continuous challenge. Our approach mixes both parallelism and specialized computer architecture. We have assembled a 48-node cluster called RDISK (Reconfigurable Devices for Intelligent Storage toward Knowledge-discovery), to perform very fast scanning of genomic databases. Each node houses a hard disk drive tightly connected to reconfigurable hardware (FPGA technology). Depending on the pattern, a custom hardware filter is mapped (in less than one second) to the hard disk output, which allows data to be processed on the fly, with relevant information then being reported.
|
Discovering Olfactory Receptors
In collaboration with CNRS UMR6061 laboratory, RDISK has been used to scan the draft genome of the dog (36 Gbp consisting of 36 million unassembled sequences) for olfactory receptors. Models created from well-known receptors were translated to automata hardwired in RDISK nodes. We found a total of 1122 olfactory receptors, including more than 400 new ones. The scan process takes ten minutes on the RDISK cluster. In comparison, it takes two days on a standard Pentium 4 (2.5 GHz) PC.
From Genes to Cells: Dynamics of Biological Networks
Most of a cell's phenotypes result from the intricate interactions of a large number of cellular constituents. Modelling the dynamics of such systems is a challenging issue, since the lack of quantitative data means a qualitative modelling approach is required. Qualitative differential models appear to be appropriate for studying genetically regulated metabolisms such as lipid metabolism: it has been proven that this system changes its functioning mode by equilibrium shifts instead of the multi-stationary equilibrium switches that usually appear in genetic networks. As a complementary approach, we study the combinatorics of incoming signals in discrete event networks. For example, TGF-_ is a transcription factor implied in the development of fibrosis, often leading to liver cancer. In collaboration with the INSERM U456 laboratory, we are identifying signals that spread through the TGF-_ signalling network and lead a cell to a given state. These two approaches are embedded in a graphical analyser tool called GARMeN.
A Stimulating Research Environment
All our collaborations benefit from the OUEST-genopole structure, part of a French Government initiative in favour of genomics and post-genomics studies. Founded in January 2002 in the west of France, it represents a network of more than fifty laboratories with experience in marine science, agri-business and human health. It offers technological platforms and is linked to a canceropole structure. Symbiose is in charge of the bioinformatics platform (http://genouest.org/).
Link:
http://www.irisa.fr/symbiose
Please contact:
Jacques Nicolas, IRISA-INRIA Rennes, France
Tel: +33 2 9984 7312
E-mail: Jacques.Nicolas![]() inria.fr
inria.fr

 This issue in
This issue in