BAIT: Bacteria - Antibiotic Interaction Tool
by Grainne Kerr
ALIFE is the name given to the discipline that studies natural life by attempting to recreate biological phenomena using computers and other artificial media. Through the study of the simplest organisms we are able to gain fundamental knowledge about the underlying mechanisms of life. BAIT (Bacteria Antibiotic Interaction Tool) was developed to provide a simulation tool for such a study. Using only simple rules governing the behaviour of individual bacteria cells with each other and with the environment population behaviour can be investigated.
Complex agent based systems consist of many similar and simple components. The system as a whole often has complex behaviour, which is more than the sum of its constituent parts. Agents can be used to conceptualise and implement such a software modelling application. The fundamental unit of bacteria life is the cell; it therefore seems appropriate to model each bacteria cell as an agent. BAIT was developed to provide a platform whereby the underlying mechanism governing bacteria growth could be investigated. Using the agent based approach the bacteria can be treated as individual entities and hence the model is more general and quantitative. Without specifying a global algorithm and using only rules at a local level of each agent, the growth patterns of bacteria will emerge. By specifying rules for bacteria interactions with the environment and obtaining the correct growth curves we can get an understanding of the underlying principals of bacteria behaviour.
Bacteria growth can be divided into four distinct phases - lag phase, exponential phase, stationary phase and death phase. How the bacteria thrive within a particular environment is largely dependant on the classification of the bacteria. Bacteria can be classified under a number of headings. A bacterium can be classified according to optimum growth temperature (Mesophile, Thermophile, psychrophile), optimum ph (acidophile, neurtophile, alkaliphile), gram staining (gram positive, gram negative), motility (high, low) etc. These factors influence how a bacteria strain will grow and survive within a particular environment eg an acidophile bacterium will not thrive in a alkaline environment, not all bacteria are motile.
The affect of an antibiotic on a particular bacterium can be classified as either bactericidal or bacteriostatic. A bactericide is a substance that has the ability to kill bacteria. In contrast bacteriostatic antibiotics, while not directly lethal to bacteria, significantly hamper their growth by interfering with processes involved in protein production, DNA production and cellular metabolism. Antibiotics can also be classified according to mode of action and effective range. Effective range refers to the range of bacteria that a particular antibiotic is effective against eg wide spread will effect both gram positive and gram-negative bacteria. Mode of action refers to the method of inhibiting the cells natural working processes. Two main types: cell wall inhibitors and protein synthesis inhibitors.
BAIT was developed using the rules derived from microbial literature. Each bacterium is represented as an individual agent, all having the same potential behaviour and conforming to the same rule principals. Associated with each bacterium is a set of parameters that model certain aspects of the agent such as temperature classification, ph classification, metabolism, motility, minimum energy required for cell maintenance etc. By setting the relevant parameters for the individual agents a range of bacteria types can be modelled. Implementing as simple rules as possible for the interactions of the agents with each other and with the environment the emergent growth patterns can be observed eg when replication stock reaches a pre defined threshold, divide.
|
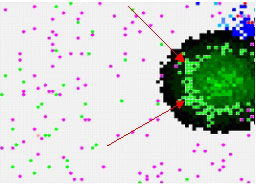
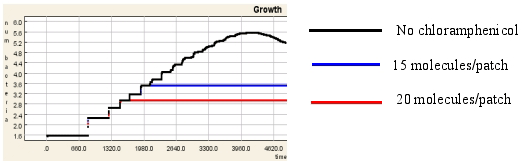
Implementing 'simple rules' for the behaviour of bacteria within an environment growth curves exhibiting the four characteristics stages of microbial growth can be observed. When antibiotic is included in the environment the effects are evident not only on the graphs generated (see Figure 1) but also visually in the virtual world (see Figure 2). In the case of the bactericidal antibiotics in particular, the bacteria manifest themselves in the form of a zone of inhibition around the diffusing mass of antibiotic molecules.
|
Using the agent based approach the four distinct phases are produced for population growth of bacteria while only specifying bacterial action at a local cell level. When the environment is altered the growth of the bacterial population responds in a manner predicted from in vitro experimentation. This simulation offers a flexible approach to viewing the effects of individual behaviour on bacterial growth and provides an excellent mechanism for understanding the behaviour of individual bacteria. Modelling the effects of antibiotic further advances the simulation. The user can localize the antibiotic in a particular area and allow it to diffuse out into the agar. Using this tool the effect of antibiotic under a wide variety of environmental conditions can the studied.
BAIT was developed at Dublin City University. Contributers are Marian Duggan, Grainne Kerr, Christopher Pender and Ronan Winters.
Please contact:
Grainne Kerr, Dublin City University/IUC, Ireland
E-mail: grainne.kerr![]() computing.dcu.ie
computing.dcu.ie

 This issue in
This issue in